Testing a PA model
Testing Problem
We are interested in the following testing problem: how do we distinguish between a graph generated from a PA model, and a graph from a configuration model with the same degree distribution as the first graph?
The point of this test is, in part, to determine what are the defining properties of a PA model besides the fact that it is scale-free. Of course, we consider the undirected version, as otherwise it would be trivial (since PA vertices all have an out-degree of \(m\)).
Our first idea was to take inspiration from (Bubeck, Devroye, and Lugosi 2014Bubeck, Sébastien, Luc Devroye, and Gábor Lugosi. 2014. “Finding Adam in random growing trees.” arXiv.org, November. http://arxiv.org/abs/1411.3317v1.), and consider centroids1 defined as the vertex whose removal results in the minimum largest component. However, given the lack of an off-the-shelf implementation of this algorithm in R, we decided to look for a simpler test. As it turns out, if we just look at the nodes with the highest degree (together), we get interesting results.
set.seed(5)
library(igraph)
# create induced subgraph by taking the k largest degree nodes
create_induced_topk <- function(g, k) {
n <- gorder(g)
dg <- degree(g)
induced_subgraph(g, order(dg)[(n-k+1):n])
}
# compares to graphs g,h to test which is which
tester <- function(g, h, k=20) {
g2 <- create_induced_topk(g, k)
h2 <- create_induced_topk(h, k)
# if sizes are the same, we should return 1/2
ifelse(gsize(g2) == gsize(h2), .5, gsize(g2) < gsize(h2))
}Consider the following test: for a graph \(G\), calculate the induced subgraph by taking the top-\(k\) degree vertices. Which model produces an induced subgraph with more edges?
n <- 100
k <- 5
g <- sample_pa(n, power=1, m=2)
(g2 <- create_induced_topk(g, k))## IGRAPH 99eb979 D--- 5 7 -- Barabasi graph
## + attr: name (g/c), power (g/n), m (g/n), zero.appeal (g/n), algorithm
## | (g/c)
## + edges from 99eb979:
## [1] 2->1 3->1 3->2 4->1 4->3 5->1 5->4h <- sample_degseq(degree(g), method="vl")
(h2 <- create_induced_topk(h, k))## IGRAPH bdfa659 U--- 5 6 -- Degree sequence random graph
## + attr: name (g/c), method (g/c)
## + edges from bdfa659:
## [1] 1--2 3--4 1--5 2--5 3--5 4--5par(mfrow=c(1,2))
Plot(g2)
Plot(h2)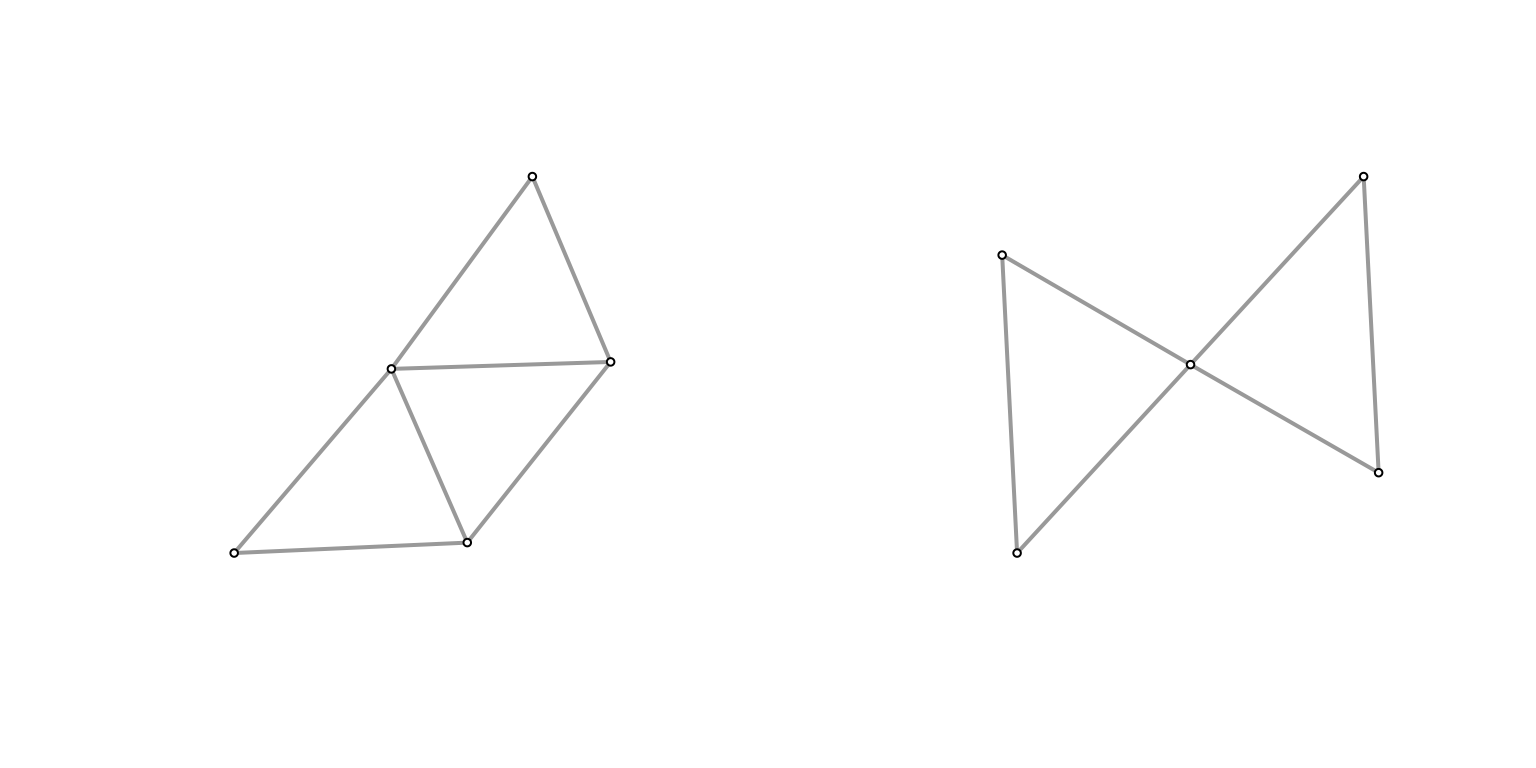
They don’t look that different at \((n,k) = (100,5)\), and the size of the graph is similar (edge count). What if we turn it up a notch?
n <- 1000
k <- 10
g <- sample_pa(n, power=1, m=2)
(g2 <- create_induced_topk(g, k))## IGRAPH fdeb1f2 D--- 10 13 -- Barabasi graph
## + attr: name (g/c), power (g/n), m (g/n), zero.appeal (g/n), algorithm
## | (g/c)
## + edges from fdeb1f2:
## [1] 2->1 3->1 3->2 4->1 4->3 5->1 6->2 7->4 7->6 8->1 9->3 9->6
## [13] 10->3h <- sample_degseq(degree(g), method="vl")
(h2 <- create_induced_topk(h, k))## IGRAPH 1468645 U--- 10 22 -- Degree sequence random graph
## + attr: name (g/c), method (g/c)
## + edges from 1468645:
## [1] 1-- 3 2-- 3 1-- 4 2-- 4 3-- 4 1-- 5 2-- 5 1-- 6 2-- 6 4-- 6 3-- 7 4-- 7
## [13] 1-- 8 3-- 8 6-- 8 7-- 8 1-- 9 3-- 9 1--10 3--10 4--10 5--10par(mfrow=c(1,2))
Plot(g2)
Plot(h2)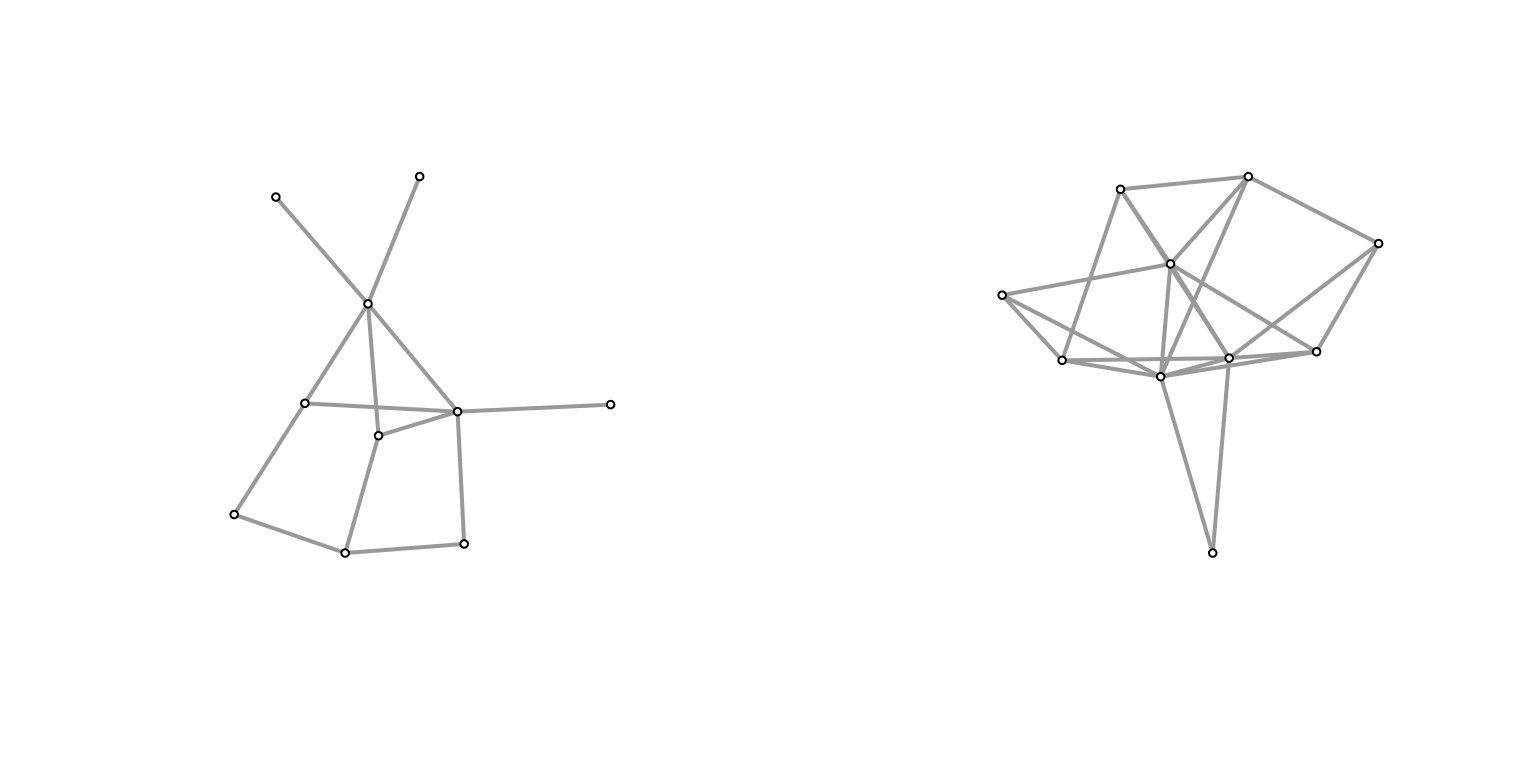
It feels like the induced subgraph from the configuration model is more densely connected (closer to a clique). Why might this be? In the case of the configuration model, if you consider the top \(k\) degree’d vertices, then there’s a very high probability of them being connected to one another2 if we take the half-edge formulation, then for any half-edge, it’s more likely to be connected to those with higher degrees (since they possess more half-edges). but they themselves have more half-edges, so it should amount to more edges between them.. Meanwhile, for a PA model, roughly we have that high degree nodes are older/earlier, and so probably have a slightly larger chance of connecting to each other than, say, a younger and an older node (since early on there aren’t many other options), but this is probably a much smaller effect.
To test this, let’s run some simulations. Here, I’ve run all combinations of \(n \in [100, 500, 1000]\) and \(k \in [5, 10, 15]\).
nsim <- 1000
ns <- c(100, 500, 1000)
ks <- c(5, 10, 20)
df <- expand.grid(k=ks, n=ns)
df$prob <- 0
cnt <- 1
for (k in ks) {
for (n in ns) {
res <- c()
for (i in 1:nsim) {
# generate graph
g <- sample_pa(n, power=1, m=2)
# generate the equivalent configuration model
h <- sample_degseq(degree(g), method="vl")
res <- c(res, tester(g, h, k=k))
}
df$prob[cnt] <- mean(res)
cnt <- cnt + 1
}
}
knitr::kable(df)| k | n | prob |
|---|---|---|
| 5 | 100 | 0.7475 |
| 10 | 100 | 0.8180 |
| 20 | 100 | 0.8705 |
| 5 | 500 | 0.9740 |
| 10 | 500 | 0.9940 |
| 20 | 500 | 0.9980 |
| 5 | 1000 | 0.9990 |
| 10 | 1000 | 1.0000 |
| 20 | 1000 | 1.0000 |
Clearly, we see a positive relationship between both \(k\) and \(n\) with respect to testing accuracy. Though, at \(n=1000\), it’s essentially as good as it’s going to get.
“Theory”
Here’s some hand-wavy calculations. For the simplest case of \(k=2\), we are interested in the probability that the top two degree vertices are connected. Let \(d_1, d_2\) be these degrees (and the respective vertex too). For the configuration model, for any given half-edge of \(d_1\), the probability that it’ll be connected to \(d_2\) is given by \(\dfrac{d_2}{D - d_1}\). Then, the probability that they’re connected is given, roughly, by \(\dfrac{d_1 \cdot d_2}{D}\).
Now consider the preferential attachment model. For now, let’s assume that \(d_1\) arrived before \(d_2\).3 can’t really WLOG here, but it feels close enough In other words, what we’re interested is the probability that one of the \(m\) vertices chosen by \(d_2\) was \(d_1\). Denote the arrival time4 which also corresponds to the number of vertices at that time of these two vertices as \(t_1\) and \(t_2\), respectively. Then, the probability is given by \(\dfrac{d_1(t_2)}{m \cdot t_2}\). The further out the \(t_2\), the larger the denominator, but also the more time \(d_1\) has had to amass its popularity.
The key is that it’s independent of \(d_2\) (i.e. the degree of the second vertex) – it would be the same probability for any other node at time \(t_2\).